21 Comparison Report: GC/FL
- Cancer Category: lymphomas
- Chip hu35ksuba: 1953 filtered probes
- Chip hu6800: NA filtered probes
Overall Complexity & Entropy Assessment
| Measure | All Filtered Probes | GO MF Terms | KEGG Pathways | HALLMARK Genes |
|---|---|---|---|---|
| Complexity | strong loss (uncertain) | () | strong loss (uncertain) | strong loss (uncertain) |
| Shannon Entropy | no clear change (uncertain) | () | mildly chaotic (well supported) | no clear change (well supported) |
| Spectral Entropy | no clear change (uncertain) | () | no clear change (uncertain) | no clear change (uncertain) |
Notes - Gain = increased complexity from normal to tumor
- Loss = complexity reduction from normal to tumor
- Chaotic = increased entropy from normal to tumor
- Anti-Chaotic = entropy reduction from normal to tumor
- All p-values unadjusted unless noted
⚠️ Table not available.
⚠️ Table not available.
Significant GO Molecular Function Terms (Complexity)
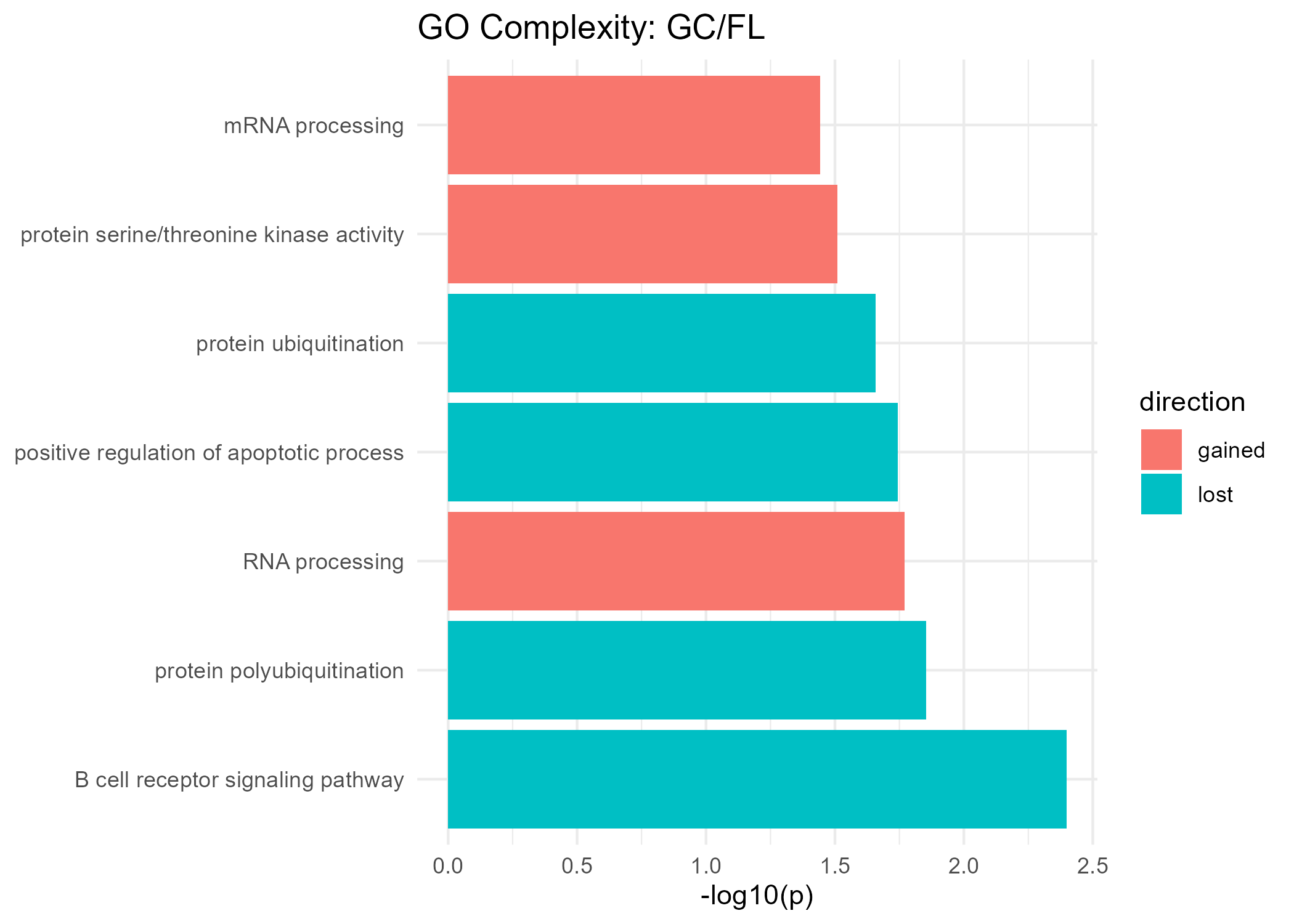
Gained Complexity
| GeneSet | p-value |
|---|---|
| RNA processing | 0.017 |
| mRNA processing | 0.036 |
| protein serine/threonine kinase activity | 0.031 |
Lost Complexity
| GeneSet | p-value |
|---|---|
| B cell receptor signaling pathway | 0.004 |
| positive regulation of apoptotic process | 0.018 |
| protein polyubiquitination | 0.014 |
| protein ubiquitination | 0.022 |
Significant GO Molecular Function Terms (Spectral Entropy)
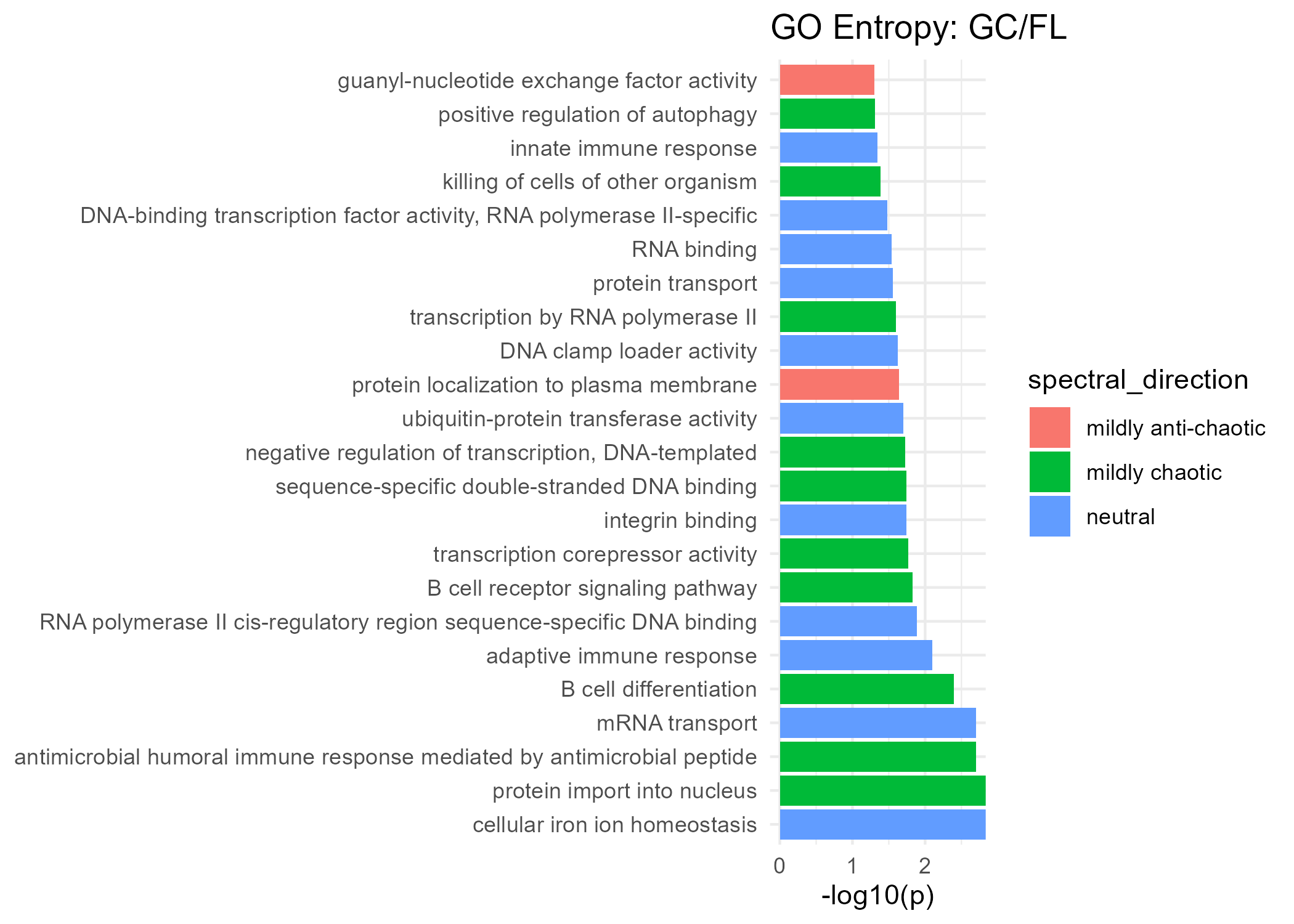
Mildly Anti-Chaotic
| GeneSet | p-value |
|---|---|
| guanyl-nucleotide exchange factor activity | 0.050 |
| protein localization to plasma membrane | 0.023 |
Neutral
| GeneSet | p-value |
|---|---|
| DNA clamp loader activity | 0.024 |
| DNA-binding transcription factor activity, RNA polymerase II-specific | 0.033 |
| RNA binding | 0.029 |
| RNA polymerase II cis-regulatory region sequence-specific DNA binding | 0.013 |
| adaptive immune response | 0.008 |
| cellular iron ion homeostasis | 0.000 |
| innate immune response | 0.045 |
| integrin binding | 0.018 |
| mRNA transport | 0.002 |
| protein transport | 0.028 |
| ubiquitin-protein transferase activity | 0.020 |
Mildly Chaotic
| GeneSet | p-value |
|---|---|
| B cell differentiation | 0.004 |
| B cell receptor signaling pathway | 0.015 |
| antimicrobial humoral immune response mediated by antimicrobial peptide | 0.002 |
| killing of cells of other organism | 0.041 |
| negative regulation of transcription, DNA-templated | 0.019 |
| positive regulation of autophagy | 0.049 |
| protein import into nucleus | 0.000 |
| sequence-specific double-stranded DNA binding | 0.018 |
| transcription by RNA polymerase II | 0.025 |
| transcription corepressor activity | 0.017 |
Significant KEGG Pathways (Complexity)
⚠️ Plot not available.Lost Complexity
| GeneSet | p-value | Cancer Pathway |
|---|---|---|
| Pathways in cancer | 0.392 | ✅ |
Significant KEGG Pathways (Spectral Entropy)
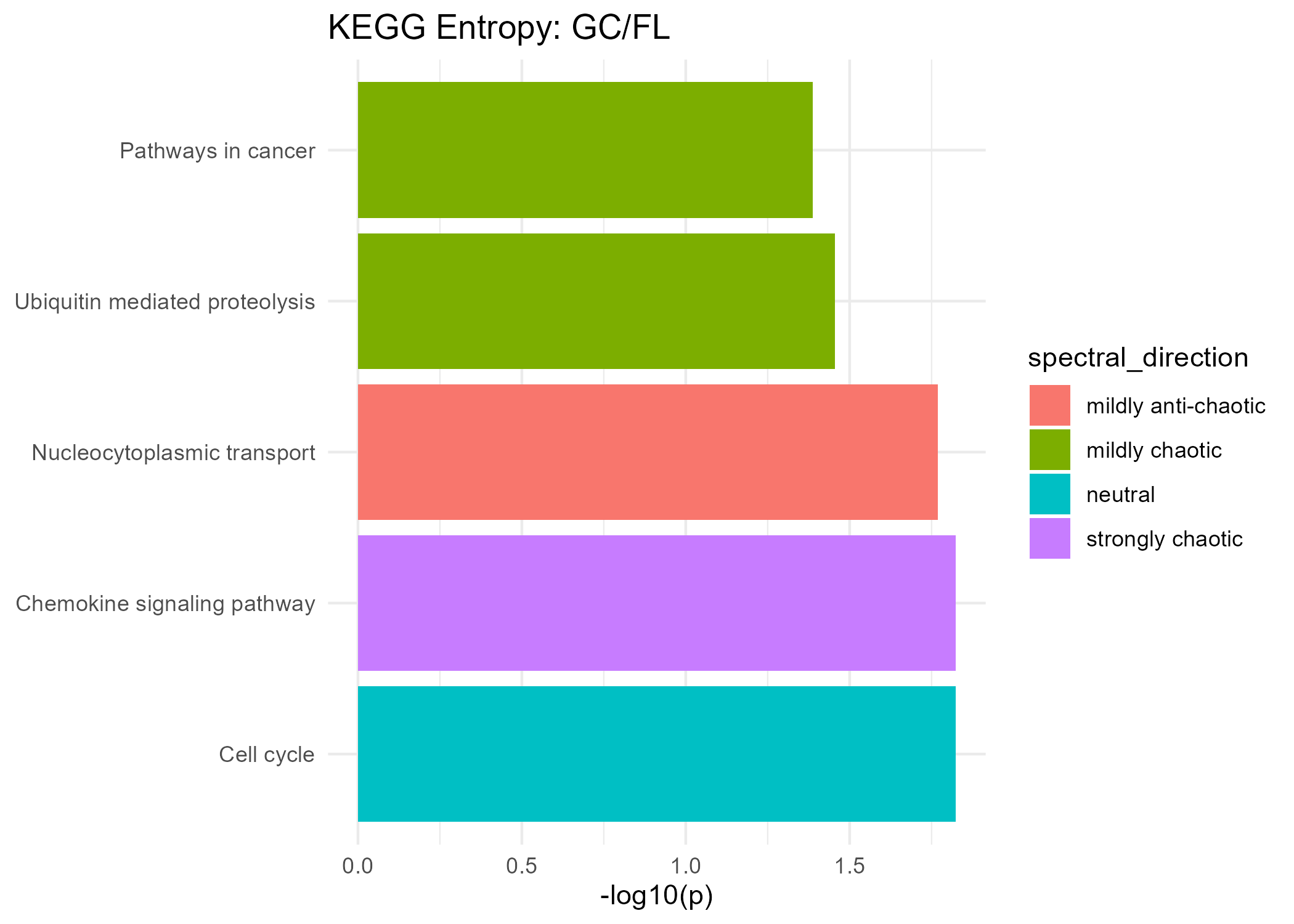
Mildly Anti-Chaotic
| GeneSet | p-value | Cancer Pathway |
|---|---|---|
| Nucleocytoplasmic transport | 0.017 |
Neutral
| GeneSet | p-value | Cancer Pathway |
|---|---|---|
| Cell cycle | 0.015 |
Mildly Chaotic
| GeneSet | p-value | Cancer Pathway |
|---|---|---|
| Pathways in cancer | 0.041 | ✅ |
| Ubiquitin mediated proteolysis | 0.035 |
Strongly Chaotic
| GeneSet | p-value | Cancer Pathway |
|---|---|---|
| Chemokine signaling pathway | 0.015 |
Significant HALLMARK Genes (Complexity)
No significant MSigDB gene sets found.
Significant HALLMARK Genes (Spectral Entropy)
Neutral
| GeneSet | p-value |
|---|---|
| HALLMARK_UV_RESPONSE_DN | 0.025 |