15 Comparison Report: LU/LUAD
- Cancer Category: carcinomas
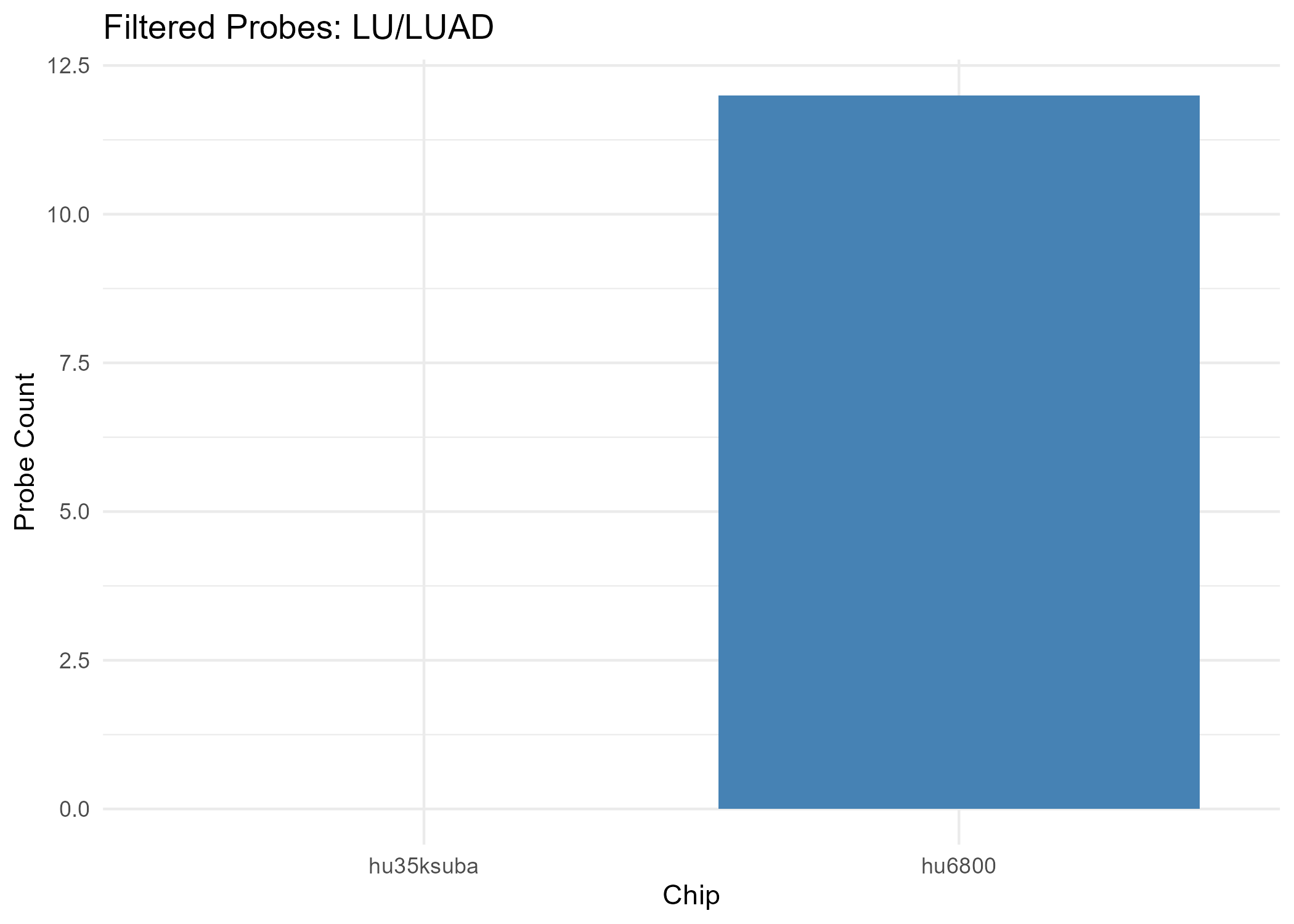
- Chip hu35ksuba: 0 filtered probes
- Chip hu6800: 64 filtered probes
Overall Complexity & Entropy Assessment
| Measure | All Filtered Probes | GO MF Terms | KEGG Pathways | HALLMARK Genes |
|---|---|---|---|---|
| Complexity | strong gain (uncertain) | () | strong gain (uncertain) | () |
| Shannon Entropy | no clear change (uncertain) | () | no clear change (uncertain) | () |
| Spectral Entropy | no clear change (uncertain) | () | no clear change (uncertain) | () |
Notes - Gain = increased complexity from normal to tumor
- Loss = complexity reduction from normal to tumor
- Chaotic = increased entropy from normal to tumor
- Anti-Chaotic = entropy reduction from normal to tumor
- All p-values unadjusted unless noted
No significant GO clusters found.
⚠️ Table not available.
Significant GO Molecular Function Terms (Complexity)
⚠️ Plot not available.No significant GO terms found.
Significant GO Molecular Function Terms (Spectral Entropy)
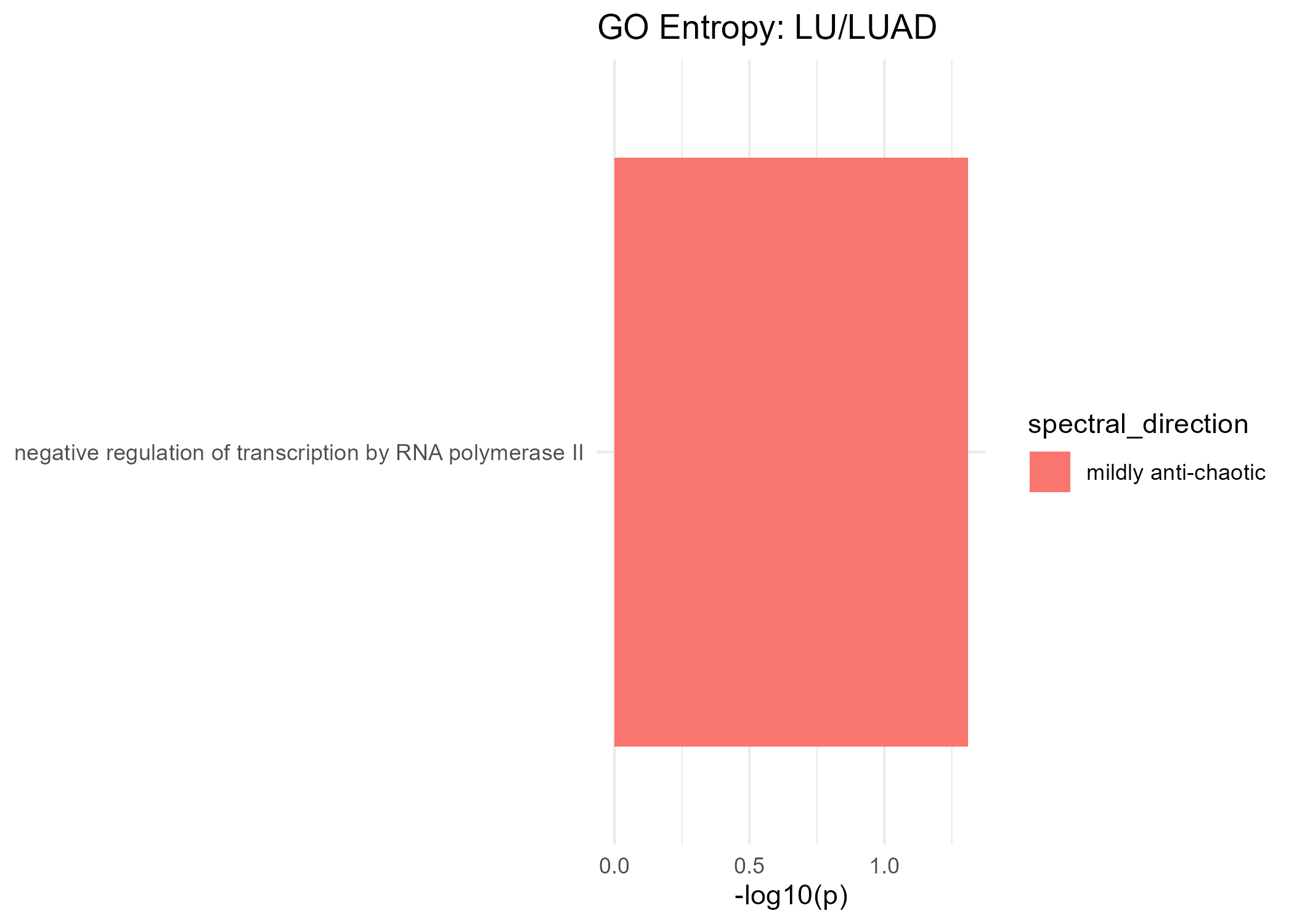
Mildly Anti-Chaotic
| GeneSet | p-value |
|---|---|
| negative regulation of transcription by RNA polymerase II | 0.049 |
Significant KEGG Pathways (Complexity)
⚠️ Plot not available.No significant KEGG pathways found (complexity).
Significant KEGG Pathways (Spectral Entropy)
⚠️ Plot not available.No significant KEGG pathways found (spectral entropy).
Significant HALLMARK Genes (Complexity)
No significant MSigDB gene sets found.
Significant HALLMARK Genes (Spectral Entropy)
No significant MSigDB gene sets found (spectral entropy).