24 Comparison Report: PB/T-ALL
- Cancer Category: leukemias
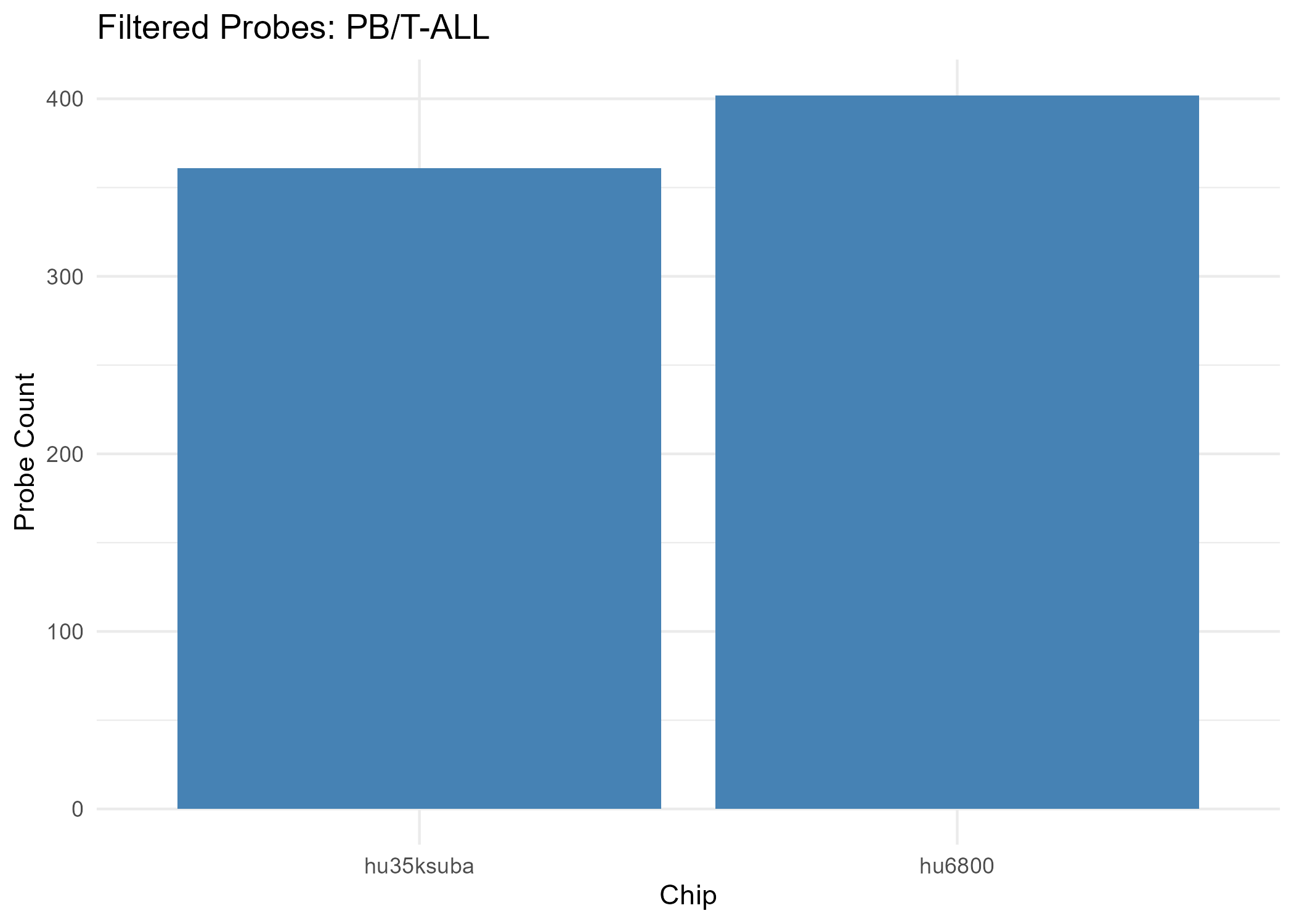
- Chip hu35ksuba: 1600 filtered probes
- Chip hu6800: 994 filtered probes
Overall Complexity & Entropy Assessment
| Measure | All Filtered Probes | GO MF Terms | KEGG Pathways | HALLMARK Genes |
|---|---|---|---|---|
| Complexity | mild gain (uncertain) | () | strong gain (uncertain) | strong gain (moderately supported) |
| Shannon Entropy | no clear change (uncertain) | () | strongly chaotic (uncertain) | mildly chaotic (uncertain) |
| Spectral Entropy | no clear change (uncertain) | () | strongly anti-chaotic (well supported) | strongly anti-chaotic (moderately supported) |
Notes - Gain = increased complexity from normal to tumor
- Loss = complexity reduction from normal to tumor
- Chaotic = increased entropy from normal to tumor
- Anti-Chaotic = entropy reduction from normal to tumor
- All p-values unadjusted unless noted
⚠️ Table not available.
⚠️ Table not available.
Significant GO Molecular Function Terms (Complexity)
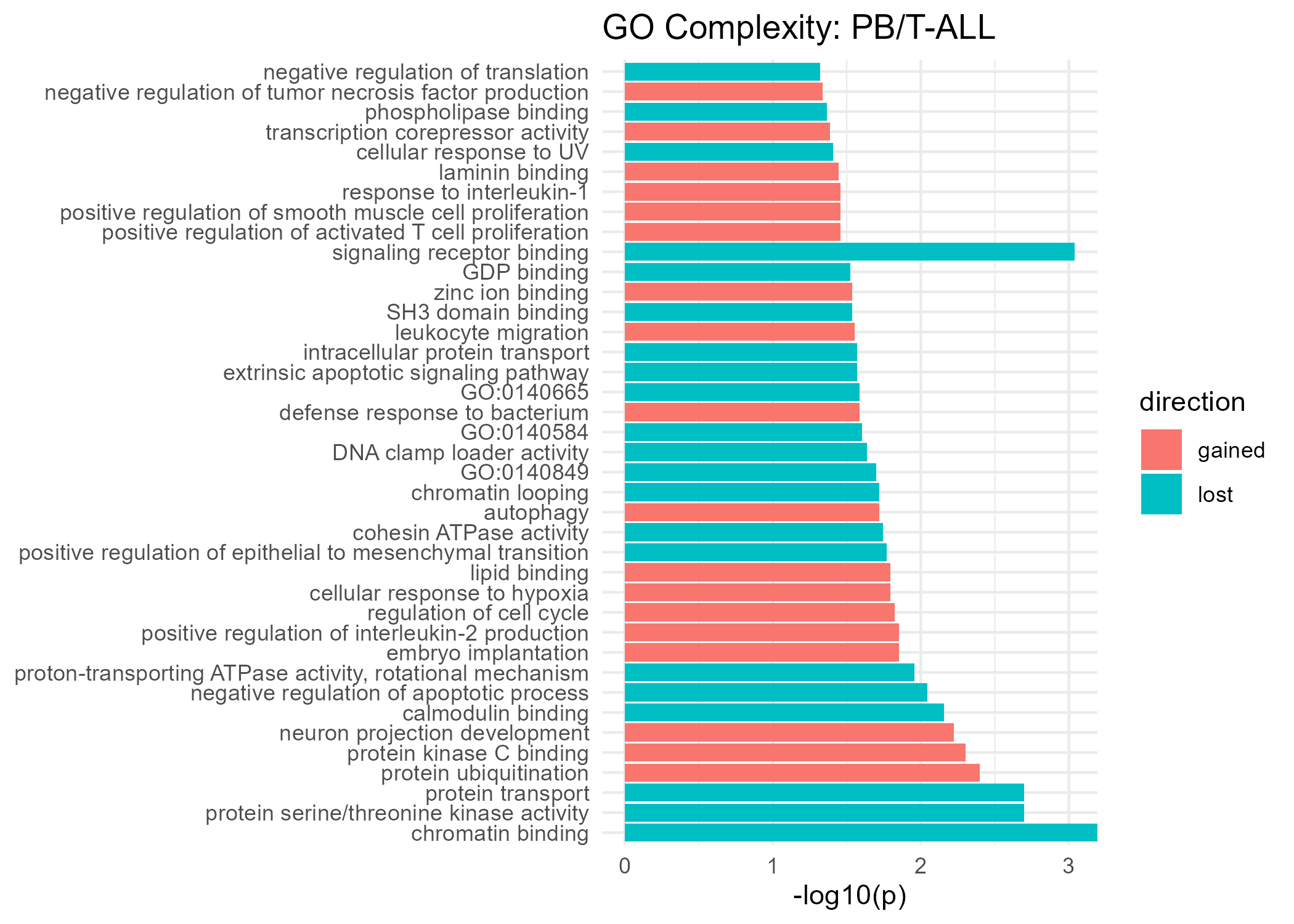
Gained Complexity
| GeneSet | p-value |
|---|---|
| autophagy | 0.019 |
| cellular response to hypoxia | 0.016 |
| defense response to bacterium | 0.026 |
| embryo implantation | 0.014 |
| laminin binding | 0.036 |
| leukocyte migration | 0.028 |
| lipid binding | 0.016 |
| negative regulation of tumor necrosis factor production | 0.046 |
| neuron projection development | 0.006 |
| positive regulation of activated T cell proliferation | 0.035 |
| positive regulation of interleukin-2 production | 0.014 |
| positive regulation of smooth muscle cell proliferation | 0.035 |
| protein kinase C binding | 0.005 |
| protein ubiquitination | 0.004 |
| regulation of cell cycle | 0.015 |
| response to interleukin-1 | 0.035 |
| transcription corepressor activity | 0.041 |
| zinc ion binding | 0.029 |
Lost Complexity
| GeneSet | p-value |
|---|---|
| DNA clamp loader activity | 0.023 |
| GDP binding | 0.030 |
| GO:0140584 | 0.025 |
| GO:0140665 | 0.026 |
| GO:0140849 | 0.020 |
| SH3 domain binding | 0.029 |
| calmodulin binding | 0.007 |
| cellular response to UV | 0.039 |
| chromatin binding | 0.000 |
| chromatin looping | 0.019 |
| cohesin ATPase activity | 0.018 |
| extrinsic apoptotic signaling pathway | 0.027 |
| intracellular protein transport | 0.027 |
| negative regulation of apoptotic process | 0.009 |
| negative regulation of translation | 0.048 |
| phospholipase binding | 0.043 |
| positive regulation of epithelial to mesenchymal transition | 0.017 |
| protein serine/threonine kinase activity | 0.002 |
| protein transport | 0.002 |
| proton-transporting ATPase activity, rotational mechanism | 0.011 |
| signaling receptor binding | 0.026 |
| signaling receptor binding | 0.035 |
Significant GO Molecular Function Terms (Spectral Entropy)
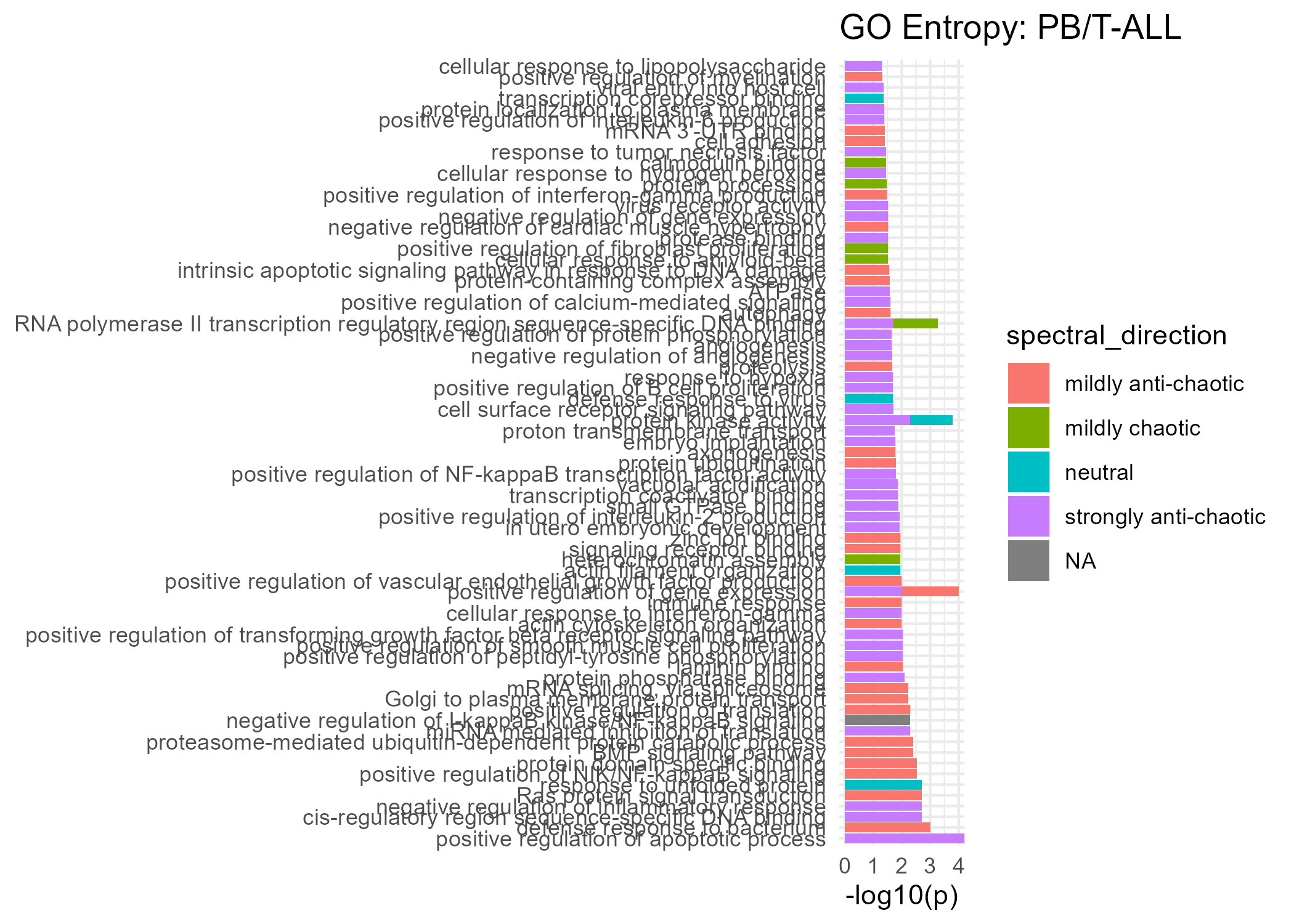
Strongly Anti-Chaotic
| GeneSet | p-value |
|---|---|
| ATPase | 0.026 |
| RNA polymerase II transcription regulatory region sequence-specific DNA binding | 0.019 |
| angiogenesis | 0.023 |
| cell surface receptor signaling pathway | 0.019 |
| cellular response to hydrogen peroxide | 0.035 |
| cellular response to interferon-gamma | 0.010 |
| cellular response to lipopolysaccharide | 0.049 |
| cis-regulatory region sequence-specific DNA binding | 0.002 |
| embryo implantation | 0.017 |
| in utero embryonic development | 0.012 |
| miRNA mediated inhibition of translation | 0.005 |
| negative regulation of angiogenesis | 0.022 |
| negative regulation of gene expression | 0.031 |
| negative regulation of inflammatory response | 0.002 |
| positive regulation of B cell proliferation | 0.020 |
| positive regulation of NF-kappaB transcription factor activity | 0.016 |
| positive regulation of apoptotic process | 0.000 |
| positive regulation of calcium-mediated signaling | 0.025 |
| positive regulation of gene expression | 0.010 |
| positive regulation of interleukin-2 production | 0.012 |
| positive regulation of interleukin-6 production | 0.041 |
| positive regulation of peptidyl-tyrosine phosphorylation | 0.009 |
| positive regulation of protein phosphorylation | 0.023 |
| positive regulation of smooth muscle cell proliferation | 0.009 |
| positive regulation of transforming growth factor beta receptor signaling pathway | 0.009 |
| protease binding | 0.030 |
| protein kinase activity | 0.005 |
| protein localization to plasma membrane | 0.041 |
| protein phosphatase binding | 0.008 |
| proton transmembrane transport | 0.018 |
| response to hypoxia | 0.020 |
| response to tumor necrosis factor | 0.036 |
| small GTPase binding | 0.013 |
| transcription coactivator binding | 0.014 |
| vacuolar acidification | 0.014 |
| viral entry into host cell | 0.044 |
| virus receptor activity | 0.031 |
Mildly Anti-Chaotic
| GeneSet | p-value |
|---|---|
| BMP signaling pathway | 0.004 |
| Golgi to plasma membrane protein transport | 0.006 |
| Ras protein signal transduction | 0.002 |
| actin cytoskeleton organization | 0.010 |
| autophagy | 0.025 |
| axonogenesis | 0.017 |
| cell adhesion | 0.039 |
| defense response to bacterium | 0.001 |
| immune response | 0.010 |
| intrinsic apoptotic signaling pathway in response to DNA damage | 0.027 |
| laminin binding | 0.009 |
| mRNA 3’-UTR binding | 0.039 |
| mRNA splicing, via spliceosome | 0.006 |
| negative regulation of cardiac muscle hypertrophy | 0.031 |
| positive regulation of NIK/NF-kappaB signaling | 0.003 |
| positive regulation of gene expression | 0.010 |
| positive regulation of interferon-gamma production | 0.033 |
| positive regulation of myelination | 0.048 |
| positive regulation of translation | 0.005 |
| positive regulation of vascular endothelial growth factor production | 0.010 |
| proteasome-mediated ubiquitin-dependent protein catabolic process | 0.004 |
| protein domain specific binding | 0.003 |
| protein ubiquitination | 0.016 |
| protein-containing complex assembly | 0.026 |
| proteolysis | 0.021 |
| signaling receptor binding | 0.011 |
| zinc ion binding | 0.011 |
Neutral
| GeneSet | p-value |
|---|---|
| actin filament organization | 0.011 |
| defense response to virus | 0.020 |
| protein kinase activity | 0.032 |
| response to unfolded protein | 0.002 |
| transcription corepressor binding | 0.044 |
Mildly Chaotic
| GeneSet | p-value |
|---|---|
| RNA polymerase II transcription regulatory region sequence-specific DNA binding | 0.028 |
| calmodulin binding | 0.036 |
| cellular response to amyloid-beta | 0.030 |
| heterochromatin assembly | 0.011 |
| positive regulation of fibroblast proliferation | 0.030 |
| protein processing | 0.033 |
Uncategorized
| GeneSet | p-value |
|---|---|
| negative regulation of I-kappaB kinase/NF-kappaB signaling | 0.005 |
Significant KEGG Pathways (Complexity)
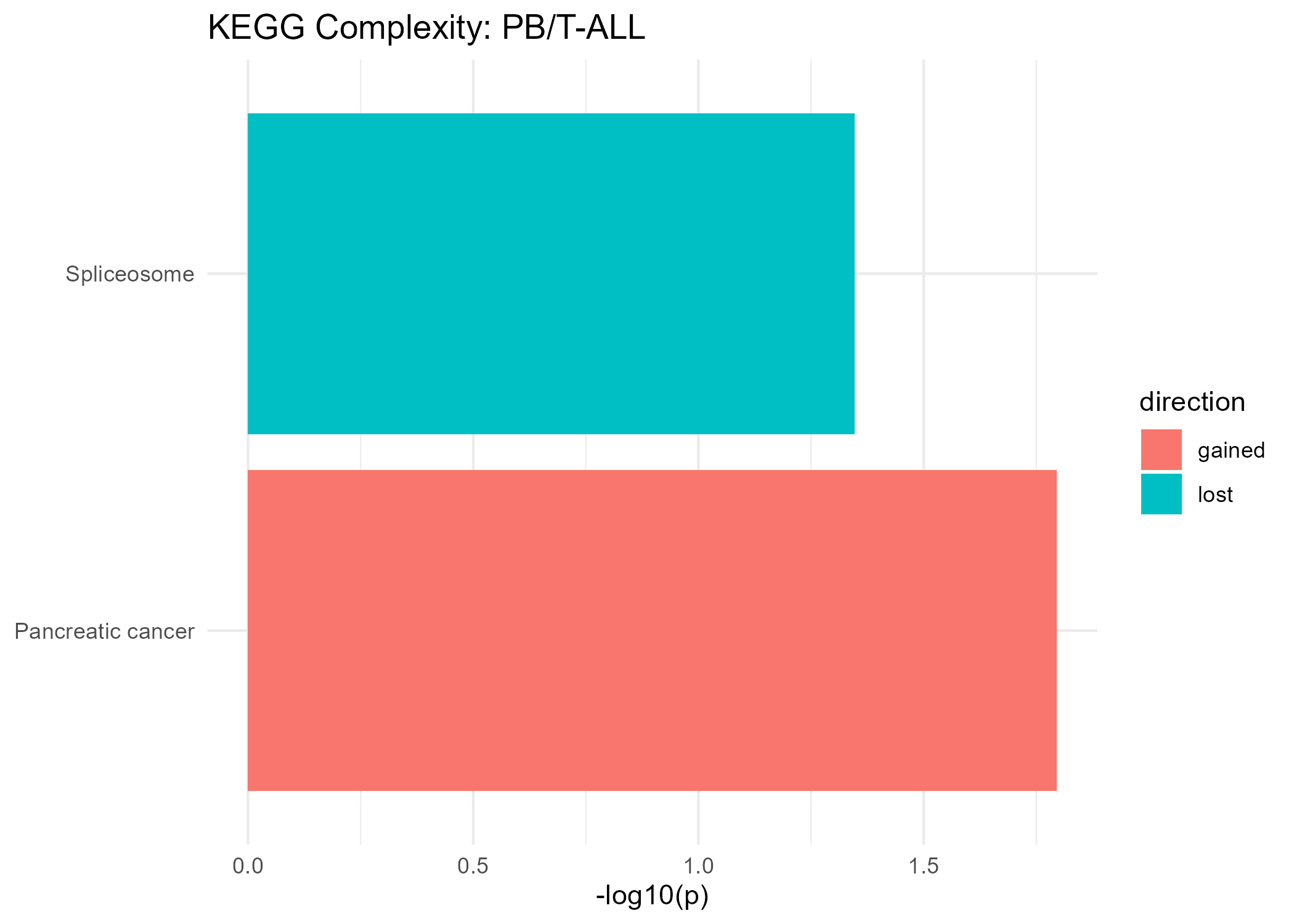
Gained Complexity
| GeneSet | p-value | Cancer Pathway |
|---|---|---|
| Pancreatic cancer | 0.016 | ✅ |
| Pathways in cancer | 0.153 | ✅ |
Lost Complexity
| GeneSet | p-value | Cancer Pathway |
|---|---|---|
| Pathways in cancer | 0.144 | ✅ |
| Prostate cancer | 0.105 | ✅ |
| Spliceosome | 0.045 |
Significant KEGG Pathways (Spectral Entropy)
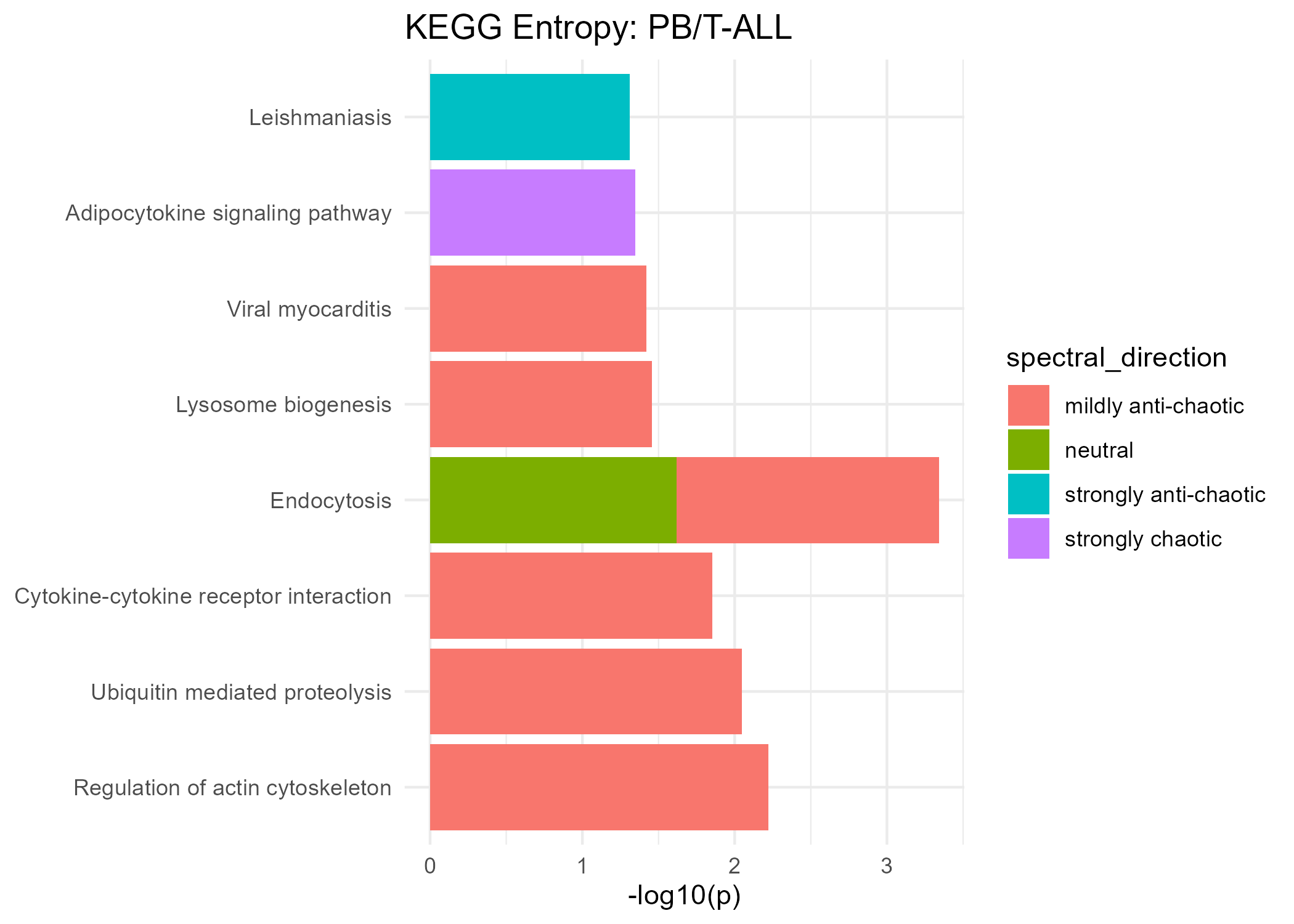
Strongly Anti-Chaotic
| GeneSet | p-value | Cancer Pathway |
|---|---|---|
| Leishmaniasis | 0.049 | |
| Pathways in cancer | 0.345 | ✅ |
| Pathways in cancer | 1.000 | ✅ |
Mildly Anti-Chaotic
| GeneSet | p-value | Cancer Pathway |
|---|---|---|
| Cytokine-cytokine receptor interaction | 0.014 | |
| Endocytosis | 0.019 | |
| Lysosome biogenesis | 0.035 | |
| Pancreatic cancer | 0.697 | ✅ |
| Prostate cancer | 0.202 | ✅ |
| Regulation of actin cytoskeleton | 0.006 | |
| Ubiquitin mediated proteolysis | 0.009 | |
| Viral myocarditis | 0.038 |
Neutral
| GeneSet | p-value | Cancer Pathway |
|---|---|---|
| Endocytosis | 0.024 |
Strongly Chaotic
| GeneSet | p-value | Cancer Pathway |
|---|---|---|
| Adipocytokine signaling pathway | 0.045 |
Significant HALLMARK Genes (Complexity)
Gained Complexity
| GeneSet | p-value |
|---|---|
| HALLMARK_EPITHELIAL_MESENCHYMAL_TRANSITION | 0.043 |
| HALLMARK_MTORC1_SIGNALING | 0.049 |
Significant HALLMARK Genes (Spectral Entropy)
Mildly Anti-Chaotic
| GeneSet | p-value |
|---|---|
| HALLMARK_ESTROGEN_RESPONSE_EARLY | 0.010 |
| HALLMARK_IL2_STAT5_SIGNALING | 0.005 |